By Megan Molteni for WIRED.
Patient Number Two was born to first-time parents, late 20s, white. The pregnancy was normal and the birth uncomplicated. But after a few months, it became clear something was wrong. The child had ear infection after ear infection and trouble breathing at night. He was small for his age, and by his fifth birthday, still hadn’t spoken. He started having seizures. Brain MRIs, molecular analyses, basic genetic testing, scores of doctors; nothing turned up answers. With no further options, in 2015 his family decided to sequence their exomes — the portion of the genome that codes for proteins — to see if he had inherited a genetic disorder from his parents. A single variant showed up: ARID1B.
The mutation suggested he had a disease called Coffin-Siris syndrome. But Patient Number Two didn’t have that disease’s typical symptoms, like sparse scalp hair and incomplete pinky fingers. So, doctors, including Karen Gripp, who met with Two’s family to discuss the exome results, hadn’t really considered it. Gripp was doubly surprised when she uploaded a photo of Two’s face to Face2Gene. The app, developed by the same programmers who taught Facebook to find your face in your friend’s photos, conducted millions of tiny calculations in rapid succession—how much slant in the eye? How narrow is that eyelid fissure? How low are the ears? Quantified, computed, and ranked to suggest the most probable syndromes associated with the facial phenotype. There’s even a heat map overlay on the photo that shows which the features are the most indicative match.
“In hindsight it was all clear to me,” says Gripp, who is chief of the Division of Medical Genetics at A.I. duPont Hospital for Children in Delaware, and had been seeing the patient for years. “But it hadn’t been clear to anyone before.” What had taken Patient Number Two’s doctors 16 years to find took Face2Gene just a few minutes.
“What had taken Patient Number Two’s doctors 16 years to find took Face2Gene just a few minutes.”
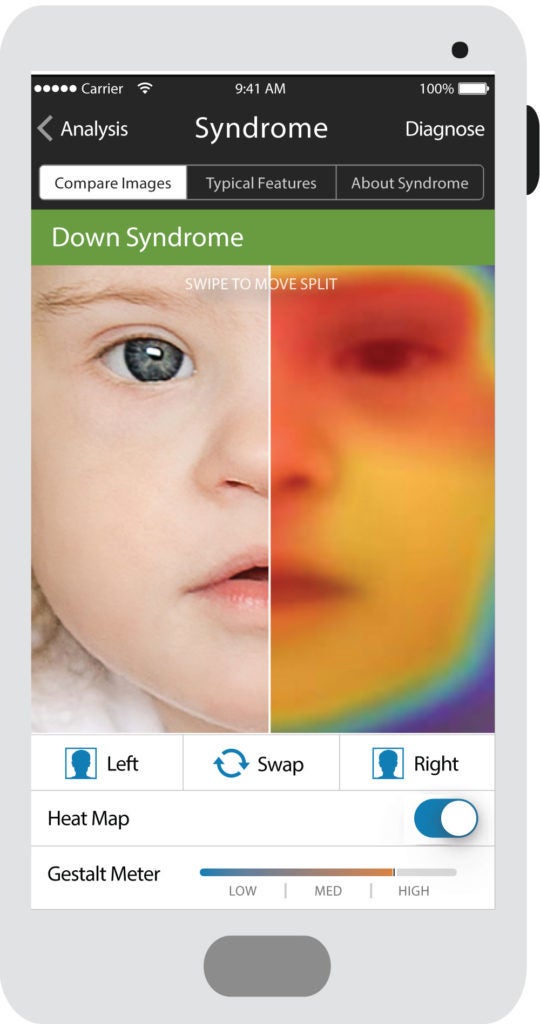
Face2Gene takes advantage of the fact that so many genetic conditions have a tell-tale “face”—a unique constellation of features that can provide clues to a potential diagnosis. It is just one of several new technologies taking advantage of how quickly modern computers can analyze, sort, and find patterns across huge reams of data. They are built in fields of artificial intelligence known as deep learning and neural nets — among the most promising to deliver AI’s 50-year old promise to revolutionize medicine by recognizing and diagnosing disease.
Genetic syndromes aren’t the only diagnoses that could get help from machine learning. The RightEye GeoPref Autism Test can identify the early stages of autism in infants as young as 12 months — the crucial stages where early intervention can make a big difference. Unveiled January 2 at CES in Las Vegas, the technology uses infrared sensors test the child’s eye movement as they watch a split-screen video: one side fills with people and faces, the other with moving geometric shapes. Children at that age should be much more attracted to faces than abstract objects, so the amount of time they look at each screen can indicate where on the autism spectrum a child might fall.
In validation studies done by the test’s inventor, UC San Diego researcher Karen Pierce,1 the test correctly predicted autism spectrum disorder 86 percent of the time in more than 400 toddlers. That said, it’s still pretty new, and hasn’t yet been approved by the FDA as a diagnostic tool. “In terms of machine learning, it’s the simplest test we have,” says RightEye’s Chief Science Officer Melissa Hunfalvay. “But before this, it was just physician or parent observations that might lead to a diagnosis. And the problem with that is it hasn’t been quantifiable.”
A similar tool could help with early detection of America’s sixth leading cause of death: Alzheimer’s disease. Often, doctors don’t recognize physical symptoms in time to try any of the disease’s few existing interventions. But machine learning hears what doctor’s can’t: Signs of cognitive impairment in speech. This is how Toronto-based Winterlight Labs is developing a tool to pick out hints of dementia in its very early stages. Co-founder Frank Rudzicz calls these clues “jitters,” and “shimmers:” high frequency wavelets only computers, not humans, can hear.
Winterlight’s tool is way more sensitive than the pencil and paper-based tests doctor’s currently use to assess Alzheimer’s. Besides being crude, data-wise, those tests can’t be taken more than once every six months. Rudzicz’s tool can be used multiple times a week, which lets it track good days, bad days, and measure a patient’s cognitive functions over time. The product is still in beta, but is currently being piloted by medical professionals in Canada, the US, and France.
If this all feels a little scarily sci-fi to you, it’s useful to remember that doctors have been trusting computers with your diagnoses for a long time. That’s because machines are much more sensitive at both detecting and analyzing the many subtle indications that our bodies are misbehaving. For instance, without computers, Patient Number Two would never have been able to compare his exome to thousands of others, and find the genetic mutation marking him with Coffin-Siris syndrome.
But none of this makes doctors obsolete. Even Face2Gene — which, according to its inventors, can diagnose up to half of the 8,000 known genetic syndromes using facial patterns gleaned from the hundreds of thousands of images in its database — needs a doctor (like Karen Gripp) with enough experience to verify the results. In that way, machines are an extension of what medicine has always been: A science that grows more powerful with every new data point.
More from Wired:

